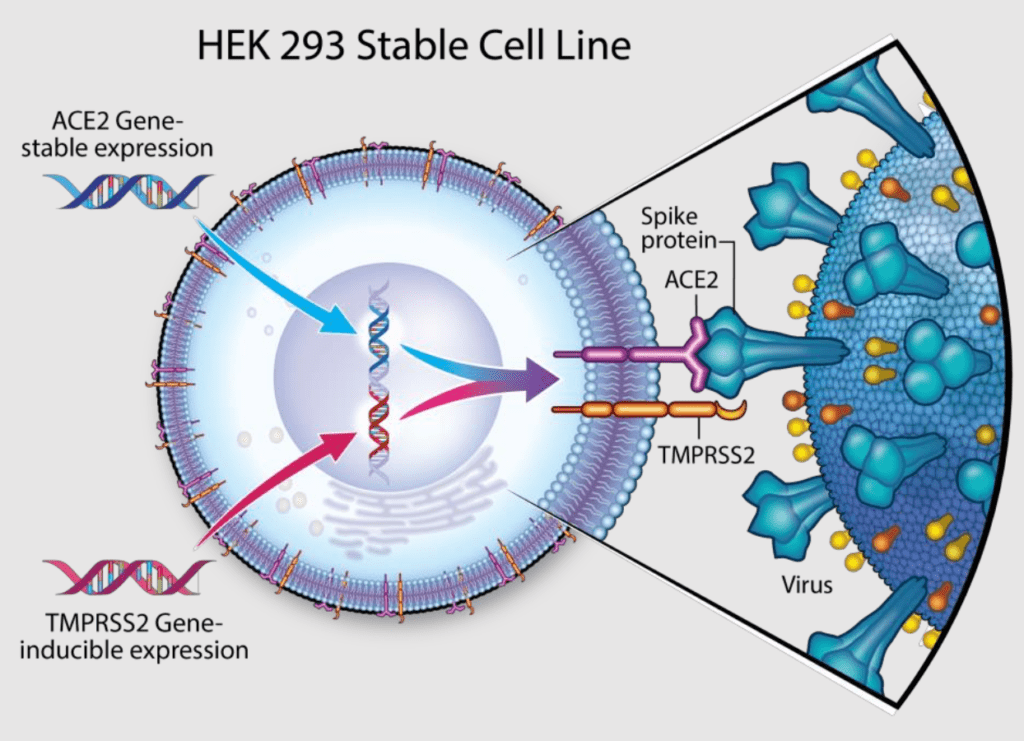
Blocking viral entry into host cells using antibodies and small molecules has been an effective antiviral strategy. Specifically to SARs, the spike proteins expressed on the surface of SARS-CoV and SARS-CoV-2 bind to angiotensin-converting enzyme 2 (ACE2) on the host cell surface. They are primed by serine protease TMPRSS2 which inturn induces the viral entry into host cells.
To develop therapeutic agents that can block SARS-CoV-2 viral entry, it is essential to measure their effectiveness in interfering protein-protein interaction or protease enzymatic activity. To screen for and optimize compounds with these properties, Multispan created a SARS-CoV-2 pseudo-viral entry assay in vitro. The system is in high throughput format that can be conducted in any standard Biosafety Level 1 (BSL1) lab.