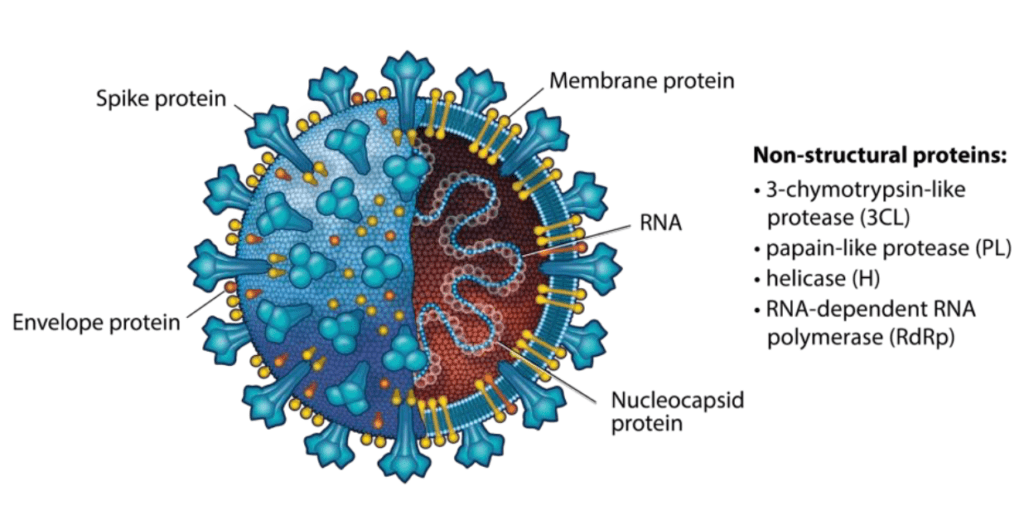
Rapid Repurposing
Cell-based Assay: RdRp Inhibitor Screening
Repurposing Beyond Remdesivir
Cell-based Assay: Protease
Blocking SARS From the Start
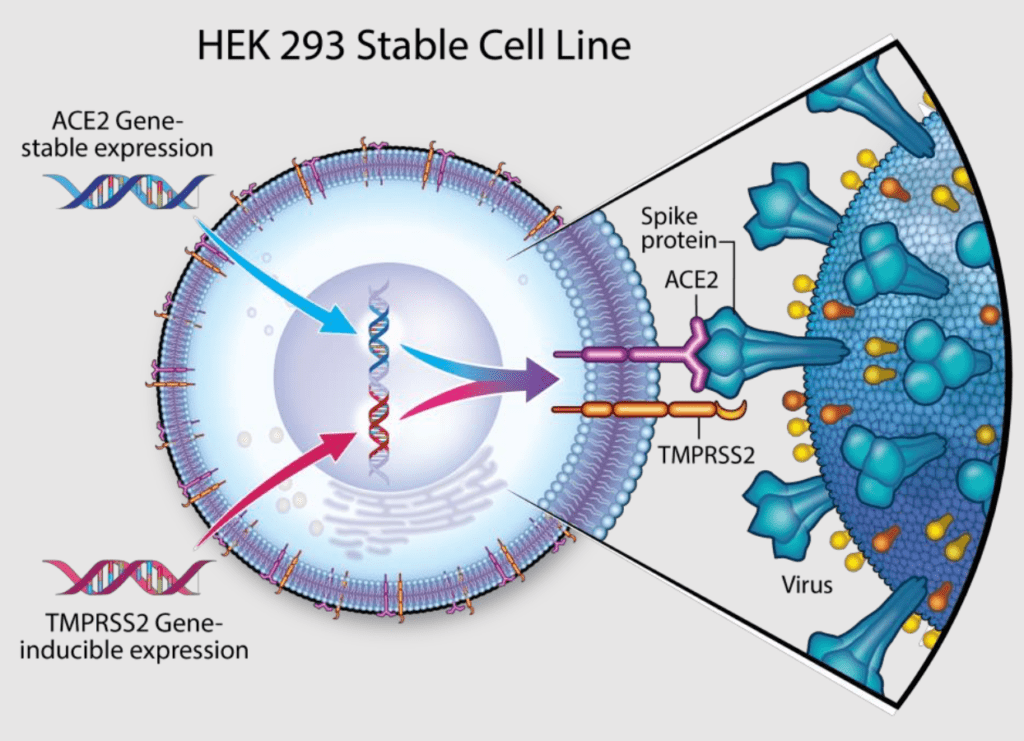
Cell-based Assay: Viral Entry
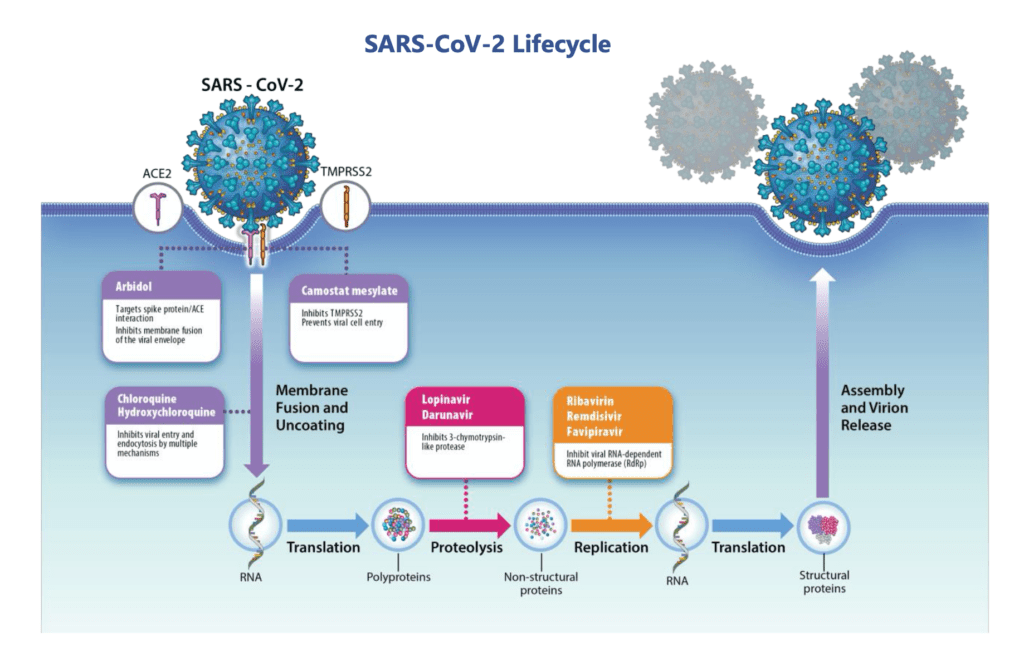
Lessons Learned From HIV
- Use a cocktail of drugs
- Target multiple independent viral pathways or enzymes
- Continue to develop drugs with incremental improvements in potency, efficacy, and safety profiles
Helping Make a Difference
References:
1. Mulangu, et al. A Randomized, Controlled Trial of Ebola Virus Disease Therapeutics. (2019) N. Engl. J. Med., 381(24):2293-2303.
2. Wang, et.al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. (2020) Cell Res., 30:269-71.
3. Beigel, et al. Remdesivir for the Treatment of Covid-19 – Preliminary Report. (2020) N. Engl. J. Med., 2020 May 22:NEJMoa2007764. doi: 10.1056/NEJMoa2007764. Online ahead of print.
4. Adaptive COVID-19 Treatment Trial 2 (ACTT-II):https://clinicaltrials.gov/ct2/show/NCT04401579
5. Domling and Gao. Chemistry and Biology of SARS-CoV-2, (2020) Chem., https://doi.org/10.1016/j.chempr.2020.04.02
6. Zhang, et al. Angiotensin- converting enzyme 2 (ACE2) as a SARS- CoV-2 receptor: molecular mechanisms and potential therapeutic target. (2020). Intensive Care Med., 46, 586–5905.
7. Hoffmann, et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. (2020) Cell, 181, 271–280.
8. Pipeline Review. APEIRON’s respiratory drug product to start pilot clinical trial to treat coronavirus disease COVID-19 in China. Pipeline Review https://pipelinereview.com/index.php/2020022673884/Proteins- and- Peptides/APEIRONs- respiratory- drugproduct-to- start- pilot- clinical- trial- to- treat- coronavirusdisease- COVID-19- in- China.html (2020).
9. Marcus, et al. Narrowing the gap in life expectancy for HIV+ compared with HIV- individuals. Conference on Retroviruses and Opportunistic Infections (CROI), Boston, abstract 54, 2016.
Other Suggested Reading:
Li, et al. Therapeutic options for the 2019 novel coronavirus (2019-nCoV). (2020) Nat. Rev. Drug Discov. 19, 149–150.
Tay, et al. The trinity of COVID-19: immunity, inflammation and intervention. Nat Rev Immunol. (2020) 20(6):363-374. doi: 10.1038/s41577-020-0311-8. Epub 2020 Apr 28.
Oberfield, et al. SnapShot: COVID-19. (2020) Cell 181, May 14, 2020 doi.org/10.1016/j.cell.2020.04.013.