Product Information
Catalog Number:
Cpc1104-1B
Lot Number:
Cpc1104-1B-020216
Quantity:
1 vial (2 x 106) frozen cells
Freeze Medium:
Sigma Freezing Medium (C-6164)
Host cell:
CHO K1
Transfection:
Expression vector containing full-length human FFAR2 cDNA (GenBank Accession Number XM_015441021) with FLAG tag sequence at N-terminus
Recommended Storage:
Liquid nitrogen upon receiving
Propagation Medium: DMEM/F12, 10% FBS, 10 μg/mL puromycin
Stability:
In progress
Data Sheet
Background: GPR43, or free fatty acid receptor 2 (FFAR2), encodes a deduced 330- amino acid protein with 7 transmembrane domains. GPR43 is expressed by enteroendocrine L cells containing peptide YY in the large intestine, adipocytes, and peripheral blood mononuclear cells. The GPR43 receptor binds short-chain fatty acids composed of less than 5 carbon atoms. Many short-chain fatty acids are produced by gut bacteria during dietary fiber fermentation. This ligand specificity provides a molecular link between diet, gastrointestinal bacterial metabolism, and immune and inflammatory responses. The importance of GPR43 in inflammation, glucose and lipid homeostasis, and adiposity demonstrate its potential as a therapeutic target for type 2 diabetes, obesity, and ulcerative colitis.
Application: Functional assays
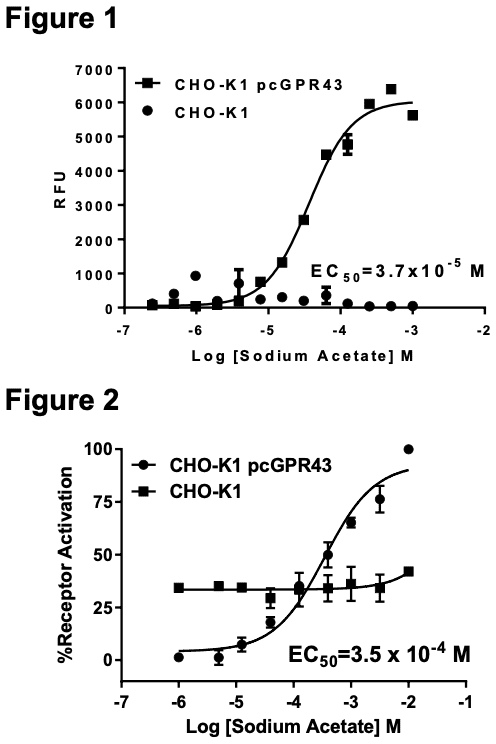
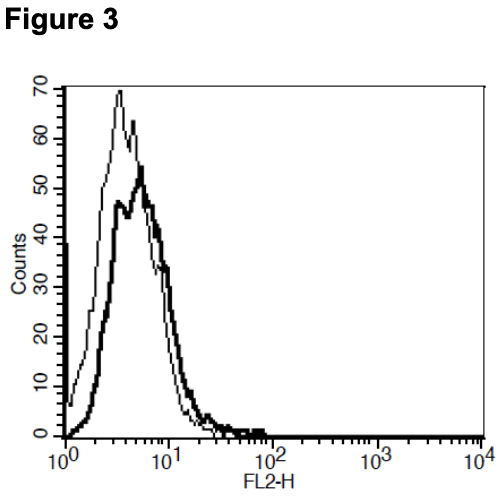
Figure 1. Dose-dependent calcium flux upon treatment with ligand, monitored with FLIPR. Figure 2. Dose-dependent inhibition of forskolin-stimulated intracellular cAMP accumulation upon treatment with ligand. Figure 3. Receptor expression on cell surface measured by flow cytometry (FACS) using an anti-FLAG antibody. Thin line: parental cells; thick line: receptor- expressing cells.
References:
Ang, Z., & Ding, J. L. (2016). GPR41 and GPR43 in Obesity and Inflammation – Protective or Causative? Frontiers in Immunology, 7, 28. http://doi.org/10.3389/fimmu.2016.00028
Sawzdargo et al. (1997) Cluster of four novel human G protein-coupled receptor genes occurring in close proximity to CD22 gene on chromosome 19q13.1. Biochem Biophys Res Commun 239:543-547.
Tikhonova, I. G., & Poerio, E. (2015). Free fatty acid receptors: structural models and elucidation of ligand binding interactions. BMC Structural Biology,15, 16. http://doi.org/10.1186/s12900-015-0044-2
